- 快召唤伙伴们来围观吧
- 微博 QQ QQ空间 贴吧
- 文档嵌入链接
- 复制
- 微信扫一扫分享
- 已成功复制到剪贴板
DataAnalytics2015 week6b
展开查看详情
1 . NY housing data revisited - Weighted kNN kmeans, clustering, some plotting, Bayes Peter Fox Data Analytics – ITWS-4963/ITWS-6965 Week 6b, March 6, 2015 1
2 . Plot tools/ tips pairs, gpairs, scatterplot.matrix, clustergram, ggmap, geocoord, etc. data() # precip, presidents, swiss, sunspot.month, environmental, ethanol, ionosphere More script fragments in Lab6b_*_2015.R on the web site (escience.rpi.edu/data/DA ) Resetting plot space: par(mfrow=c(1,1)) par()$mar # to view margins par(mai=c(0.1,0.1,0.1,0.1)) 2
3 . Do over… • Get Bronx.R from the escience script location • Look through the pdf – linked off the website for this week (6) – Data_Analytics2015_week5.pdf ~ is the week 5b lab, but working with new libraries (thanks to Jiaju and James) – skip page 1… • A lesson in R… 3
4 . Plotting clusters (DIY) library(cluster) clusplot(mapmeans, mapobj$cluster, color=TRUE, shade=TRUE, labels=2, lines=0) # Centroid Plot against 1st 2 discriminant functions #library(fpc) plotcluster(mapmeans, mapobj$cluster) • dendogram? library(fpc) • cluster.stats 4
5 . Comparing cluster fits Use help. > help(plotcluster) > help(cluster.stats) 5
6 . Using a contingency table > data(Titanic) > mdl <- naiveBayes(Survived ~ ., data = Titanic) Naive Bayes Classifier for Discrete Predictors Call: naiveBayes.formula(formula = Survived ~ ., data = Titanic) > mdl A-priori Survived probabilities: No Yes 0.676965 0.323035 Conditional probabilities: Class Survived 1st 2nd 3rd Crew No 0.08187919 0.11208054 0.35436242 0.45167785 Yes 0.28551336 0.16596343 0.25035162 0.29817159 Sex Survived Male Female No 0.91543624 0.08456376 Yes 0.51617440 0.48382560 Age Survived Child Adult 6 No 0.03489933 0.96510067 Yes 0.08016878 0.91983122 Try Lab6b_9_2014.R
7 . Classification Bayes • Retrieve the abalone.csv dataset • Predicting the age of abalone from physical measurements. • Perform naivebayes classification to get predictors for Age (Rings). Interpret. • Compare to what you got from kknn (weighted nearest neighbors) in class 4b 7
8 . http://www.ugrad.stat.ubc.ca/R/library/mlb ench/html/HouseVotes84.html require(mlbench) data(HouseVotes84) model <- naiveBayes(Class ~ ., data = HouseVotes84) predict(model, HouseVotes84[1:10,-1]) predict(model, HouseVotes84[1:10,-1], type = "raw") pred <- predict(model, HouseVotes84[,-1]) table(pred, HouseVotes84$Class) 8
9 . Hair, eye color > data(HairEyeColor) > mosaicplot(HairEyeColor) > margin.table(HairEyeColor,3) Sex Male Female 279 313 > margin.table(HairEyeColor,c(1,3)) Sex Hair Male Female Black 56 52 Brown 143 143 Red 34 37 Blond 46 81 Construct a naïve Bayes classifier and test it! 9
10 . Cluster plotting source("http://www.r-statistics.com/wp- content/uploads/2012/01/source_https.r.txt") # source code from github require(RCurl) require(colorspace) source_https("https://raw.github.com/talgalili/R-code- snippets/master/clustergram.r") data(iris) set.seed(250) par(cex.lab = 1.5, cex.main = 1.2) Data <- scale(iris[,-5]) # scaling clustergram(Data, k.range = 2:8, line.width = 0.004) #10line.width - adjust according to Y-scale
11 . Any good? set.seed(500) Data2 <- scale(iris[,-5]) par(cex.lab = 1.2, cex.main = .7) par(mfrow = c(3,2)) for(i in 1:6) clustergram(Data2, k.range = 2:8 , line.width = .004, add.center.points = T) Scale? We’ll look at multi-dimensional scaling soon (mds) 11
12 . How can you tell it is good? set.seed(250) Data <- rbind( cbind(rnorm(100,0, sd = 0.3),rnorm(100,0, sd = 0.3),rnorm(100,0, sd = 0.3)), cbind(rnorm(100,1, sd = 0.3),rnorm(100,1, sd = 0.3),rnorm(100,1, sd = 0.3)), cbind(rnorm(100,2, sd = 0.3),rnorm(100,2, sd = 0.3),rnorm(100,2, sd = 0.3))) clustergram(Data, k.range = 2:5 , line.width = .004, add.center.points = T) 12
13 . More complex… set.seed(250) Data <- rbind( cbind(rnorm(100,1, sd = 0.3),rnorm(100,0, sd = 0.3),rnorm(100,0, sd = 0.3),rnorm(100,0, sd = 0.3)), cbind(rnorm(100,0, sd = 0.3),rnorm(100,1, sd = 0.3),rnorm(100,0, sd = 0.3),rnorm(100,0, sd = 0.3)), cbind(rnorm(100,0, sd = 0.3),rnorm(100,1, sd = 0.3),rnorm(100,1, sd = 0.3),rnorm(100,0, sd = 0.3)), cbind(rnorm(100,0, sd = 0.3),rnorm(100,0, sd = 0.3),rnorm(100,0, sd = 0.3),rnorm(100,1, sd = 0.3))) clustergram(Data, k.range = 2:8 , line.width = .004, add.center.points = T) 13
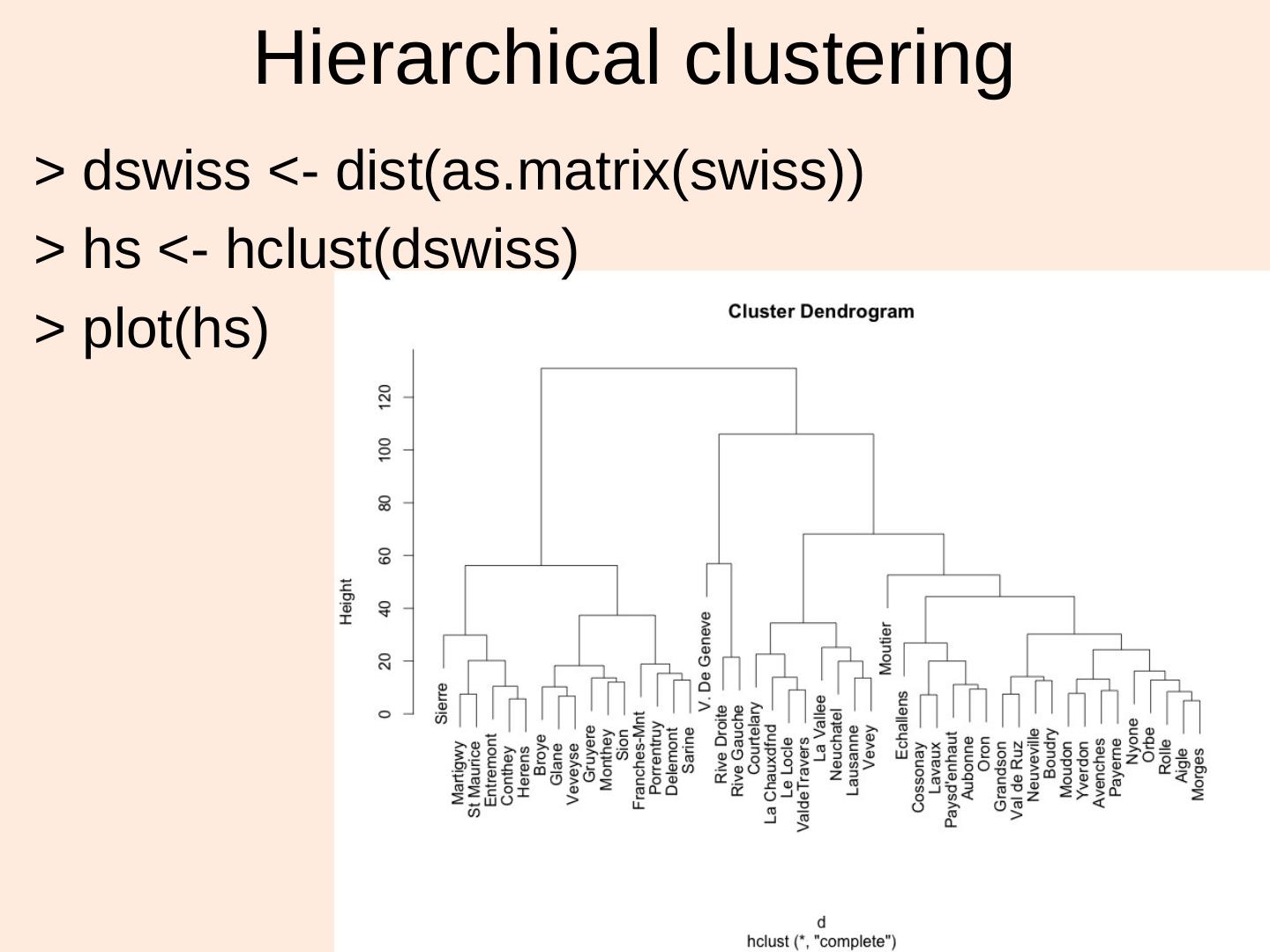
14 . Hierarchical clustering > dswiss <- dist(as.matrix(swiss)) > hs <- hclust(dswiss) > plot(hs) 14
15 . Another example > A = c(1, 2.5); B = c(5, 10); C = c(23, 34) > D = c(45, 47); E = c(4, 17); F = c(18, 4) > df <- data.frame(rbind(A,B,C,D,E,F)) > colnames(df) <- c("x","y") > hc <- hclust(dist(df)) > plot(hc) > df$cluster <- cutree(hc,k=2) # 2 clusters > plot(y~x,df,col=cluster) 15
16 . See also • Lab5a_ctree_1_2015.R – Try clustergram instead – Try hclust • Lab3b_kmeans1_2015.R – Try clustergram instead – Try hclust 16
17 . Assignment 6 preview • Your term projects should fall within the scope of a data analytics problem of the type you have worked with in class/ labs, or know of yourself – the bigger the data the better. This means that the work must go beyond just making lots of figures. You should develop the project to indicate you are thinking of and exploring the relationships and distributions within your data. Start with a hypothesis, think of a way to model and use the hypothesis, find or collect the necessary data, and do both preliminary analysis, detailed modeling and summary (interpretation). – Note: You do not have to come up with a positive result, i.e. disproving the hypothesis is just as good. Please use the section numbering below for your written submission for this assignment. • Introduction (2%) • Data Description (3%) • Analysis (8%) • Model Development (8%) • Conclusions and Discussion (4%) 17 • Oral presentation (5%) (10 mins)
18 . Assignments to come • Term project (A6). Due – early May. 30% (25% written, 5% oral; individual). Available before spring break. • Assignment 7: Predictive and Prescriptive Analytics. Due ~ week ~ 10. 15% (15% written; individual); 18